|
Protein design
Protein design is the rational design of new protein molecules to design novel activity, behavior, or purpose, and to advance basic understanding of protein function.[1] Proteins can be designed from scratch (de novo design) or by making calculated variants of a known protein structure and its sequence (termed protein redesign). Rational protein design approaches make protein-sequence predictions that will fold to specific structures. These predicted sequences can then be validated experimentally through methods such as peptide synthesis, site-directed mutagenesis, or artificial gene synthesis. Rational protein design dates back to the mid-1970s.[2] Recently, however, there were numerous examples of successful rational design of water-soluble and even transmembrane peptides and proteins, in part due to a better understanding of different factors contributing to protein structure stability and development of better computational methods. Overview and historyThe goal in rational protein design is to predict amino acid sequences that will fold to a specific protein structure. Although the number of possible protein sequences is vast, growing exponentially with the size of the protein chain, only a subset of them will fold reliably and quickly to one native state. Protein design involves identifying novel sequences within this subset. The native state of a protein is the conformational free energy minimum for the chain. Thus, protein design is the search for sequences that have the chosen structure as a free energy minimum. In a sense, it is the reverse of protein structure prediction. In design, a tertiary structure is specified, and a sequence that will fold to it is identified. Hence, it is also termed inverse folding. Protein design is then an optimization problem: using some scoring criteria, an optimized sequence that will fold to the desired structure is chosen. When the first proteins were rationally designed during the 1970s and 1980s, the sequence for these was optimized manually based on analyses of other known proteins, the sequence composition, amino acid charges, and the geometry of the desired structure.[2] The first designed proteins are attributed to Bernd Gutte, who designed a reduced version of a known catalyst, bovine ribonuclease, and tertiary structures consisting of beta-sheets and alpha-helices, including a binder of DDT. Urry and colleagues later designed elastin-like fibrous peptides based on rules on sequence composition. Richardson and coworkers designed a 79-residue protein with no sequence homology to a known protein.[2] In the 1990s, the advent of powerful computers, libraries of amino acid conformations, and force fields developed mainly for molecular dynamics simulations enabled the development of structure-based computational protein design tools. Following the development of these computational tools, great success has been achieved over the last 30 years in protein design. The first protein successfully designed completely de novo was done by Stephen Mayo and coworkers in 1997,[3] and, shortly after, in 1999 Peter S. Kim and coworkers designed dimers, trimers, and tetramers of unnatural right-handed coiled coils.[4][5] In 2003, David Baker's laboratory designed a full protein to a fold never seen before in nature.[6] Later, in 2008, Baker's group computationally designed enzymes for two different reactions.[7] In 2010, one of the most powerful broadly neutralizing antibodies was isolated from patient serum using a computationally designed protein probe.[8] Due to these and other successes (e.g., see examples below), protein design has become one of the most important tools available for protein engineering. There is great hope that the design of new proteins, small and large, will have uses in biomedicine and bioengineering. Underlying models of protein structure and functionProtein design programs use computer models of the molecular forces that drive proteins in in vivo environments. In order to make the problem tractable, these forces are simplified by protein design models. Although protein design programs vary greatly, they have to address four main modeling questions: What is the target structure of the design, what flexibility is allowed on the target structure, which sequences are included in the search, and which force field will be used to score sequences and structures. Target structure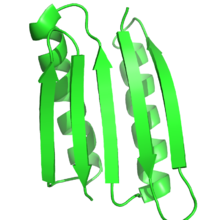 Protein function is heavily dependent on protein structure, and rational protein design uses this relationship to design function by designing proteins that have a target structure or fold. Thus, by definition, in rational protein design the target structure or ensemble of structures must be known beforehand. This contrasts with other forms of protein engineering, such as directed evolution, where a variety of methods are used to find proteins that achieve a specific function, and with protein structure prediction where the sequence is known, but the structure is unknown. Most often, the target structure is based on a known structure of another protein. However, novel folds not seen in nature have been made increasingly possible. Peter S. Kim and coworkers designed trimers and tetramers of unnatural coiled coils, which had not been seen before in nature.[4][5] The protein Top7, developed in David Baker's lab, was designed completely using protein design algorithms, to a completely novel fold.[6] More recently, Baker and coworkers developed a series of principles to design ideal globular-protein structures based on protein folding funnels that bridge between secondary structure prediction and tertiary structures. These principles, which build on both protein structure prediction and protein design, were used to design five different novel protein topologies.[9] Sequence space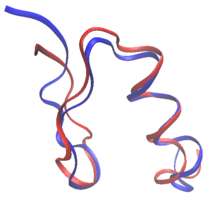 In rational protein design, proteins can be redesigned from the sequence and structure of a known protein, or completely from scratch in de novo protein design. In protein redesign, most of the residues in the sequence are maintained as their wild-type amino-acid while a few are allowed to mutate. In de novo design, the entire sequence is designed anew, based on no prior sequence. Both de novo designs and protein redesigns can establish rules on the sequence space: the specific amino acids that are allowed at each mutable residue position. For example, the composition of the surface of the RSC3 probe to select HIV-broadly neutralizing antibodies was restricted based on evolutionary data and charge balancing. Many of the earliest attempts on protein design were heavily based on empiric rules on the sequence space.[2] Moreover, the design of fibrous proteins usually follows strict rules on the sequence space. Collagen-based designed proteins, for example, are often composed of Gly-Pro-X repeating patterns.[2] The advent of computational techniques allows designing proteins with no human intervention in sequence selection.[3] Structural flexibility In protein design, the target structure (or structures) of the protein are known. However, a rational protein design approach must model some flexibility on the target structure in order to increase the number of sequences that can be designed for that structure and to minimize the chance of a sequence folding to a different structure. For example, in a protein redesign of one small amino acid (such as alanine) in the tightly packed core of a protein, very few mutants would be predicted by a rational design approach to fold to the target structure, if the surrounding side-chains are not allowed to be repacked. Thus, an essential parameter of any design process is the amount of flexibility allowed for both the side-chains and the backbone. In the simplest models, the protein backbone is kept rigid while some of the protein side-chains are allowed to change conformations. However, side-chains can have many degrees of freedom in their bond lengths, bond angles, and χ dihedral angles. To simplify this space, protein design methods use rotamer libraries that assume ideal values for bond lengths and bond angles, while restricting χ dihedral angles to a few frequently observed low-energy conformations termed rotamers. Rotamer libraries are derived from the statistical analysis of many protein structures. Backbone-independent rotamer libraries describe all rotamers.[10] Backbone-dependent rotamer libraries, in contrast, describe the rotamers as how likely they are to appear depending on the protein backbone arrangement around the side chain.[11] Most protein design programs use one conformation (e.g., the modal value for rotamer dihedrals in space) or several points in the region described by the rotamer; the OSPREY protein design program, in contrast, models the entire continuous region.[12] Although rational protein design must preserve the general backbone fold a protein, allowing some backbone flexibility can significantly increase the number of sequences that fold to the structure while maintaining the general fold of the protein.[13] Backbone flexibility is especially important in protein redesign because sequence mutations often result in small changes to the backbone structure. Moreover, backbone flexibility can be essential for more advanced applications of protein design, such as binding prediction and enzyme design. Some models of protein design backbone flexibility include small and continuous global backbone movements, discrete backbone samples around the target fold, backrub motions, and protein loop flexibility.[13][14] Energy function Rational protein design techniques must be able to discriminate sequences that will be stable under the target fold from those that would prefer other low-energy competing states. Thus, protein design requires accurate energy functions that can rank and score sequences by how well they fold to the target structure. At the same time, however, these energy functions must consider the computational challenges behind protein design. One of the most challenging requirements for successful design is an energy function that is both accurate and simple for computational calculations. The most accurate energy functions are those based on quantum mechanical simulations. However, such simulations are too slow and typically impractical for protein design. Instead, many protein design algorithms use either physics-based energy functions adapted from molecular mechanics simulation programs, knowledge based energy-functions, or a hybrid mix of both. The trend has been toward using more physics-based potential energy functions.[15] Physics-based energy functions, such as AMBER and CHARMM, are typically derived from quantum mechanical simulations, and experimental data from thermodynamics, crystallography, and spectroscopy.[16] These energy functions typically simplify physical energy function and make them pairwise decomposable, meaning that the total energy of a protein conformation can be calculated by adding the pairwise energy between each atom pair, which makes them attractive for optimization algorithms. Physics-based energy functions typically model an attractive-repulsive Lennard-Jones term between atoms and a pairwise electrostatics coulombic term[17] between non-bonded atoms.  Statistical potentials, in contrast to physics-based potentials, have the advantage of being fast to compute, of accounting implicitly of complex effects and being less sensitive to small changes in the protein structure.[19] These energy functions are based on deriving energy values from frequency of appearance on a structural database. Protein design, however, has requirements that can sometimes be limited in molecular mechanics force-fields. Molecular mechanics force-fields, which have been used mostly in molecular dynamics simulations, are optimized for the simulation of single sequences, but protein design searches through many conformations of many sequences. Thus, molecular mechanics force-fields must be tailored for protein design. In practice, protein design energy functions often incorporate both statistical terms and physics-based terms. For example, the Rosetta energy function, one of the most-used energy functions, incorporates physics-based energy terms originating in the CHARMM energy function, and statistical energy terms, such as rotamer probability and knowledge-based electrostatics. Typically, energy functions are highly customized between laboratories, and specifically tailored for every design.[16] Challenges for effective design energy functionsWater makes up most of the molecules surrounding proteins and is the main driver of protein structure. Thus, modeling the interaction between water and protein is vital in protein design. The number of water molecules that interact with a protein at any given time is huge and each one has a large number of degrees of freedom and interaction partners. Instead, protein design programs model most of such water molecules as a continuum, modeling both the hydrophobic effect and solvation polarization.[16] Individual water molecules can sometimes have a crucial structural role in the core of proteins, and in protein–protein or protein–ligand interactions. Failing to model such waters can result in mispredictions of the optimal sequence of a protein–protein interface. As an alternative, water molecules can be added to rotamers. [16]
As an optimization problem The goal of protein design is to find a protein sequence that will fold to a target structure. A protein design algorithm must, thus, search all the conformations of each sequence, with respect to the target fold, and rank sequences according to the lowest-energy conformation of each one, as determined by the protein design energy function. Thus, a typical input to the protein design algorithm is the target fold, the sequence space, the structural flexibility, and the energy function, while the output is one or more sequences that are predicted to fold stably to the target structure. The number of candidate protein sequences, however, grows exponentially with the number of protein residues; for example, there are 20100 protein sequences of length 100. Furthermore, even if amino acid side-chain conformations are limited to a few rotamers (see Structural flexibility), this results in an exponential number of conformations for each sequence. Thus, in our 100 residue protein, and assuming that each amino acid has exactly 10 rotamers, a search algorithm that searches this space will have to search over 200100 protein conformations. The most common energy functions can be decomposed into pairwise terms between rotamers and amino acid types, which casts the problem as a combinatorial one, and powerful optimization algorithms can be used to solve it. In those cases, the total energy of each conformation belonging to each sequence can be formulated as a sum of individual and pairwise terms between residue positions. If a designer is interested only in the best sequence, the protein design algorithm only requires the lowest-energy conformation of the lowest-energy sequence. In these cases, the amino acid identity of each rotamer can be ignored and all rotamers belonging to different amino acids can be treated the same. Let ri be a rotamer at residue position i in the protein chain, and E(ri) the potential energy between the internal atoms of the rotamer. Let E(ri, rj) be the potential energy between ri and rotamer rj at residue position j. Then, we define the optimization problem as one of finding the conformation of minimum energy (ET):
The problem of minimizing ET is an NP-hard problem.[14][20][21] Even though the class of problems is NP-hard, in practice many instances of protein design can be solved exactly or optimized satisfactorily through heuristic methods. AlgorithmsSeveral algorithms have been developed specifically for the protein design problem. These algorithms can be divided into two broad classes: exact algorithms, such as dead-end elimination, that lack runtime guarantees but guarantee the quality of the solution; and heuristic algorithms, such as Monte Carlo, that are faster than exact algorithms but have no guarantees on the optimality of the results. Exact algorithms guarantee that the optimization process produced the optimal according to the protein design model. Thus, if the predictions of exact algorithms fail when these are experimentally validated, then the source of error can be attributed to the energy function, the allowed flexibility, the sequence space or the target structure (e.g., if it cannot be designed for).[22] Some protein design algorithms are listed below. Although these algorithms address only the most basic formulation of the protein design problem, Equation (1), when the optimization goal changes because designers introduce improvements and extensions to the protein design model, such as improvements to the structural flexibility allowed (e.g., protein backbone flexibility) or including sophisticated energy terms, many of the extensions on protein design that improve modeling are built atop these algorithms. For example, Rosetta Design incorporates sophisticated energy terms, and backbone flexibility using Monte Carlo as the underlying optimizing algorithm. OSPREY's algorithms build on the dead-end elimination algorithm and A* to incorporate continuous backbone and side-chain movements. Thus, these algorithms provide a good perspective on the different kinds of algorithms available for protein design. In 2020 scientists reported the development of an AI-based process using genome databases for evolution-based designing of novel proteins. They used deep learning to identify design-rules.[23][24] In 2022, a study reported deep learning software that can design proteins that contain prespecified functional sites.[25][26] With mathematical guaranteesDead-end eliminationThe dead-end elimination (DEE) algorithm reduces the search space of the problem iteratively by removing rotamers that can be provably shown to be not part of the global lowest energy conformation (GMEC). On each iteration, the dead-end elimination algorithm compares all possible pairs of rotamers at each residue position, and removes each rotamer r′i that can be shown to always be of higher energy than another rotamer ri and is thus not part of the GMEC: Other powerful extensions to the dead-end elimination algorithm include the pairs elimination criterion, and the generalized dead-end elimination criterion. This algorithm has also been extended to handle continuous rotamers with provable guarantees. Although the Dead-end elimination algorithm runs in polynomial time on each iteration, it cannot guarantee convergence. If, after a certain number of iterations, the dead-end elimination algorithm does not prune any more rotamers, then either rotamers have to be merged or another search algorithm must be used to search the remaining search space. In such cases, the dead-end elimination acts as a pre-filtering algorithm to reduce the search space, while other algorithms, such as A*, Monte Carlo, Linear Programming, or FASTER are used to search the remaining search space.[14] Branch and boundThe protein design conformational space can be represented as a tree, where the protein residues are ordered in an arbitrary way, and the tree branches at each of the rotamers in a residue. Branch and bound algorithms use this representation to efficiently explore the conformation tree: At each branching, branch and bound algorithms bound the conformation space and explore only the promising branches.[14][27][28] A popular search algorithm for protein design is the A* search algorithm.[14][28] A* computes a lower-bound score on each partial tree path that lower bounds (with guarantees) the energy of each of the expanded rotamers. Each partial conformation is added to a priority queue and at each iteration the partial path with the lowest lower bound is popped from the queue and expanded. The algorithm stops once a full conformation has been enumerated and guarantees that the conformation is the optimal. The A* score f in protein design consists of two parts, f=g+h. g is the exact energy of the rotamers that have already been assigned in the partial conformation. h is a lower bound on the energy of the rotamers that have not yet been assigned. Each is designed as follows, where d is the index of the last assigned residue in the partial conformation. Integer linear programmingThe problem of optimizing ET (Equation (1)) can be easily formulated as an integer linear program (ILP).[29] One of the most powerful formulations uses binary variables to represent the presence of a rotamer and edges in the final solution, and constraints the solution to have exactly one rotamer for each residue and one pairwise interaction for each pair of residues: s.t. ILP solvers, such as CPLEX, can compute the exact optimal solution for large instances of protein design problems. These solvers use a linear programming relaxation of the problem, where qi and qij are allowed to take continuous values, in combination with a branch and cut algorithm to search only a small portion of the conformation space for the optimal solution. ILP solvers have been shown to solve many instances of the side-chain placement problem.[29] Message-passing based approximations to the linear programming dualILP solvers depend on linear programming (LP) algorithms, such as the Simplex or barrier-based methods to perform the LP relaxation at each branch. These LP algorithms were developed as general-purpose optimization methods and are not optimized for the protein design problem (Equation (1)). In consequence, the LP relaxation becomes the bottleneck of ILP solvers when the problem size is large.[30] Recently, several alternatives based on message-passing algorithms have been designed specifically for the optimization of the LP relaxation of the protein design problem. These algorithms can approximate both the dual or the primal instances of the integer programming, but in order to maintain guarantees on optimality, they are most useful when used to approximate the dual of the protein design problem, because approximating the dual guarantees that no solutions are missed. Message-passing based approximations include the tree reweighted max-product message passing algorithm,[31][32] and the message passing linear programming algorithm.[33] Optimization algorithms without guaranteesMonte Carlo and simulated annealingMonte Carlo is one of the most widely used algorithms for protein design. In its simplest form, a Monte Carlo algorithm selects a residue at random, and in that residue a randomly chosen rotamer (of any amino acid) is evaluated.[21] The new energy of the protein, Enew is compared against the old energy Eold and the new rotamer is accepted with a probability of: where β is the Boltzmann constant and the temperature T can be chosen such that in the initial rounds it is high and it is slowly annealed to overcome local minima.[12] FASTERThe FASTER algorithm uses a combination of deterministic and stochastic criteria to optimize amino acid sequences. FASTER first uses DEE to eliminate rotamers that are not part of the optimal solution. Then, a series of iterative steps optimize the rotamer assignment.[34][35] Belief propagationIn belief propagation for protein design, the algorithm exchanges messages that describe the belief that each residue has about the probability of each rotamer in neighboring residues. The algorithm updates messages on every iteration and iterates until convergence or until a fixed number of iterations. Convergence is not guaranteed in protein design. The message mi→ j(rj that a residue i sends to every rotamer (rj at neighboring residue j is defined as: Both max-product and sum-product belief propagation have been used to optimize protein design. Applications and examples of designed proteinsEnzyme designThe design of new enzymes is a use of protein design with huge bioengineering and biomedical applications. In general, designing a protein structure can be different from designing an enzyme, because the design of enzymes must consider many states involved in the catalytic mechanism. However protein design is a prerequisite of de novo enzyme design because, at the very least, the design of catalysts requires a scaffold in which the catalytic mechanism can be inserted.[36] Great progress in de novo enzyme design, and redesign, was made in the first decade of the 21st century. In three major studies, David Baker and coworkers de novo designed enzymes for the retro-aldol reaction,[37] a Kemp-elimination reaction,[38] and for the Diels-Alder reaction.[39] Furthermore, Stephen Mayo and coworkers developed an iterative method to design the most efficient known enzyme for the Kemp-elimination reaction.[40] Also, in the laboratory of Bruce Donald, computational protein design was used to switch the specificity of one of the protein domains of the nonribosomal peptide synthetase that produces Gramicidin S, from its natural substrate phenylalanine to other noncognate substrates including charged amino acids; the redesigned enzymes had activities close to those of the wild-type.[41] Semi-rational designSemi-rational design is a purposeful modification method based on a certain understanding of the sequence, structure, and catalytic mechanism of enzymes. This method is between irrational design and rational design. It uses known information and means to perform evolutionary modification on the specific functions of the target enzyme. The characteristic of semi-rational design is that it does not rely solely on random mutation and screening, but combines the concept of directed evolution. It creates a library of random mutants with diverse sequences through mutagenesis, error-prone RCR, DNA recombination, and site-saturation mutagenesis. At the same time, it uses the understanding of enzymes and design principles to purposefully screen out mutants with desired characteristics. The methodology of semi-rational design emphasizes the in-depth understanding of enzymes and the control of the evolutionary process. It allows researchers to use known information to guide the evolutionary process, thereby improving efficiency and success rate. This method plays an important role in protein function modification because it can combine the advantages of irrational design and rational design, and can explore unknown space and use known knowledge for targeted modification. Semi-rational design has a wide range of applications, including but not limited to enzyme optimization, modification of drug targets, evolution of biocatalysts, etc. Through this method, researchers can more effectively improve the functional properties of proteins to meet specific biotechnology or medical needs. Although this method has high requirements for information and technology and is relatively difficult to implement, with the development of computing technology and bioinformatics, the application prospects of semi-rational design in protein engineering are becoming more and more broad.[42] Design for affinityProtein–protein interactions are involved in most biotic processes. Many of the hardest-to-treat diseases, such as Alzheimer's, many forms of cancer (e.g., TP53), and human immunodeficiency virus (HIV) infection involve protein–protein interactions. Thus, to treat such diseases, it is desirable to design protein or protein-like therapeutics that bind one of the partners of the interaction and, thus, disrupt the disease-causing interaction. This requires designing protein-therapeutics for affinity toward its partner. Protein–protein interactions can be designed using protein design algorithms because the principles that rule protein stability also rule protein–protein binding. Protein–protein interaction design, however, presents challenges not commonly present in protein design. One of the most important challenges is that, in general, the interfaces between proteins are more polar than protein cores, and binding involves a tradeoff between desolvation and hydrogen bond formation.[43] To overcome this challenge, Bruce Tidor and coworkers developed a method to improve the affinity of antibodies by focusing on electrostatic contributions. They found that, for the antibodies designed in the study, reducing the desolvation costs of the residues in the interface increased the affinity of the binding pair.[43][44][45] Scoring binding predictionsProtein design energy functions must be adapted to score binding predictions because binding involves a trade-off between the lowest-energy conformations of the free proteins (EP and EL) and the lowest-energy conformation of the bound complex (EPL):
The K* algorithm approximates the binding constant of the algorithm by including conformational entropy into the free energy calculation. The K* algorithm considers only the lowest-energy conformations of the free and bound complexes (denoted by the sets P, L, and PL) to approximate the partition functions of each complex:[14] Design for specificityThe design of protein–protein interactions must be highly specific because proteins can interact with a large number of proteins; successful design requires selective binders. Thus, protein design algorithms must be able to distinguish between on-target (or positive design) and off-target binding (or negative design).[2][43] One of the most prominent examples of design for specificity is the design of specific bZIP-binding peptides by Amy Keating and coworkers for 19 out of the 20 bZIP families; 8 of these peptides were specific for their intended partner over competing peptides.[43][46][47] Further, positive and negative design was also used by Anderson and coworkers to predict mutations in the active site of a drug target that conferred resistance to a new drug; positive design was used to maintain wild-type activity, while negative design was used to disrupt binding of the drug.[48] Recent computational redesign by Costas Maranas and coworkers was also capable of experimentally switching the cofactor specificity of Candida boidinii xylose reductase from NADPH to NADH.[49] Protein resurfacingProtein resurfacing consists of designing a protein's surface while preserving the overall fold, core, and boundary regions of the protein intact. Protein resurfacing is especially useful to alter the binding of a protein to other proteins. One of the most important applications of protein resurfacing was the design of the RSC3 probe to select broadly neutralizing HIV antibodies at the NIH Vaccine Research Center. First, residues outside of the binding interface between the gp120 HIV envelope protein and the formerly discovered b12-antibody were selected to be designed. Then, the sequence spaced was selected based on evolutionary information, solubility, similarity with the wild-type, and other considerations. Then the RosettaDesign software was used to find optimal sequences in the selected sequence space. RSC3 was later used to discover the broadly neutralizing antibody VRC01 in the serum of a long-term HIV-infected non-progressor individual.[50] Design of globular proteinsGlobular proteins are proteins that contain a hydrophobic core and a hydrophilic surface. Globular proteins often assume a stable structure, unlike fibrous proteins, which have multiple conformations. The three-dimensional structure of globular proteins is typically easier to determine through X-ray crystallography and nuclear magnetic resonance than both fibrous proteins and membrane proteins, which makes globular proteins more attractive for protein design than the other types of proteins. Most successful protein designs have involved globular proteins. Both RSD-1, and Top7 were de novo designs of globular proteins. Five more protein structures were designed, synthesized, and verified in 2012 by the Baker group. These new proteins serve no biotic function, but the structures are intended to act as building-blocks that can be expanded to incorporate functional active sites. The structures were found computationally by using new heuristics based on analyzing the connecting loops between parts of the sequence that specify secondary structures.[51] Design of membrane proteinsSeveral transmembrane proteins have been successfully designed,[52] along with many other membrane-associated peptides and proteins.[53] Recently, Costas Maranas and his coworkers developed an automated tool[54] to redesign the pore size of Outer Membrane Porin Type-F (OmpF) from E.coli to any desired sub-nm size and assembled them in membranes to perform precise angstrom scale separation. Other applicationsOne of the most desirable uses for protein design is for biosensors, proteins that will sense the presence of specific compounds. Some attempts in the design of biosensors include sensors for unnatural molecules including TNT.[55] More recently, Kuhlman and coworkers designed a biosensor of the PAK1.[56] In a sense, protein design is a subset of battery design.[further explanation needed] See also
References
Further reading
|
Portal di Ensiklopedia Dunia
![{\displaystyle \min E_{T}=\sum _{i}{\Big [}E_{i}(r_{i})+\sum _{i\neq j}E_{ij}(r_{i},r_{j}){\Big ]}\,}](https://wikimedia.org/api/rest_v1/media/math/render/svg/3332d826843218136390cef20e4ee8e3694fc477)


![{\displaystyle h=\sum _{j=d+1}^{n}[\min _{r_{j}}(E(r_{j})+\sum _{i=1}^{d}E(r_{i},r_{j})+\sum _{k=j+1}^{n}\min _{r_{k}}E(r_{j},r_{k}))]}](https://wikimedia.org/api/rest_v1/media/math/render/svg/e143d714d94f81766d65c1ab49da42eeeed08b4a)







